Brainrender: visualising brain data in 3D
By April Cashin-Garbutt
To appreciate the intricate complexities of the brain, we need to be able to visualise and interact with anatomical brain data in 3D. While advances in technology over the last decade have allowed neuroscientists to gather 3D data from the brain, up until now, most of the software tools to visualise these brain-wide data have only produced 2D slides. In addition, the existing software has been limited to a specific brain atlas or species.
Federico Claudi, PhD student in the Branco Lab, and team at SWC, have developed an open-source Python package called brainrender, which enables interactive 3D visualisation of anatomical data and allows neuroscientists to gain a more intuitive understanding of brain structures.
“Being able to visualise and interact with brain data in 3D gives you a different appreciation for what structures look like in the brain. This is especially useful for neurons that have very complicated patterns of projections. Brainrender allows us to see just how beautiful and complex the brain really is.” Federico Claudi
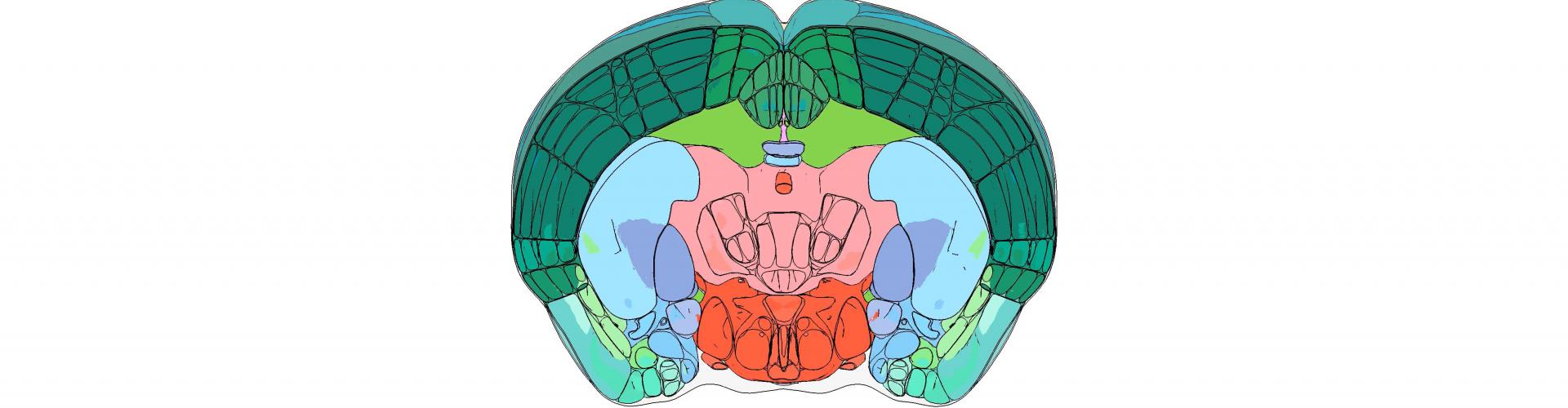
Animated video created with brainrender. Frontal view of all brain regions in the Allen Mouse Brain atlas as the brain is progressively 'sliced' in the rostro-caudal direction. Data from: https://mouse.brain-map.org/
Working with Adam Tyson, Research Fellow in the Margrie Lab at SWC, and Luigi Petrucco, PhD student at Max Planck Institute of Neurobiology in Munich, Federico and team collaborated on a project called BrainGlobe, to develop a suite of software tools for computational neuroscience. As part of this project, they developed ‘Atlas API’ which enables software, such as brainrender, to work across atlases. This means that brainrender can be used to visualise data from mice, zebrafish and other atlases without making any changes to the software itself.
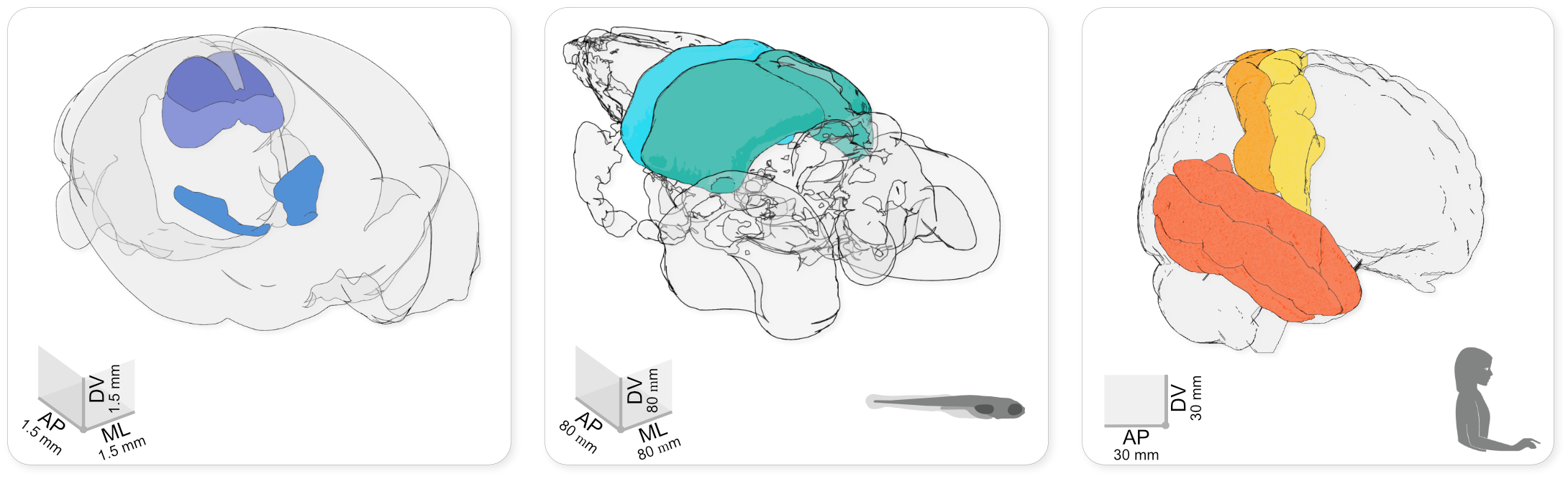
Using brainrender to visualise brain atlas data in mouse, larval zebrafish and human. The brainrender GUI. Mouse, human, and zebrafish larvae drawings from scidraw.io (doi.org/10.5281/zenodo.3925991, doi.org/10.5281/zenodo.3926189, doi.org/10.5281/zenodo.3926123).
The hope is that in addition to making brain data more intuitive to interpret, this new software will also help with dissemination and allow for more collaboration between neuroscientists working across different species. Furthermore, the software will help researchers from different labs and institutes compare data across individual species, as brainrender enables brain data to be registered to atlases.
Every brain is different and so by matching brain data to one shared template, it is easier to make comparisons. Brainrender enables the visualisation of data from different sources. For example, you can visualise your own data and alongside it, in the same renderings, you can visualise data from the Allen Institute, or the MouseLight data from Janelia, or another source. This means you can integrate information from different labs acquired with different experimental modalities.
As with all software from BrainGlobe, brainrender is open-access and can be downloaded online. There is also extensive documentation on the website so those with limited programming experience can access the tool. Federico and team also welcome users to help develop brainrender further so it can continue to benefit the neuroscience community. They hope the software can also help the development of new tools and they are currently exploring whether brainrender could assist the precise placement of electrodes and probes.
Find out more
- Read more about brainrender in eLife: Visualizing anatomically registered data with Brainrender
- Read more about BrainGlobe Atlas API in JOSS: BrainGlobe Atlas API: a common interface for neuroanatomical atlases
- Learn more about BrainGlobe: https://docs.brainglobe.info/
- Learn more about Probe planner: https://github.com/brainglobe/probeplanner