Neuroinformatics Unit paves way for open science initiatives
By Hyewon Kim
From cultural organisations painting a picture of how science can be more transparent and inclusive to collaborative computational projects that inspire rising neuroscientists around the globe, ‘open science’ is a value shared by those aiming “for the benefits of scientists and society as a whole.” Making the scientific process open and accessible to all has been the goal of many organisations in recent years.
Adam Tyson, Head of the SWC and Gatsby Computational Neuroscience Unit (GCNU) Neuroinformatics Unit (NIU), has a similar vision in mind. In spring of 2022, he returned to SWC after initially working in the Margrie Lab to form a research software engineering team. The NIU team works closely with researchers at SWC and GCNU on multiple projects spanning the enhancement of BrainGlobe, standardisation of data structures, and refinement of pipelines for microscopy images, among others.
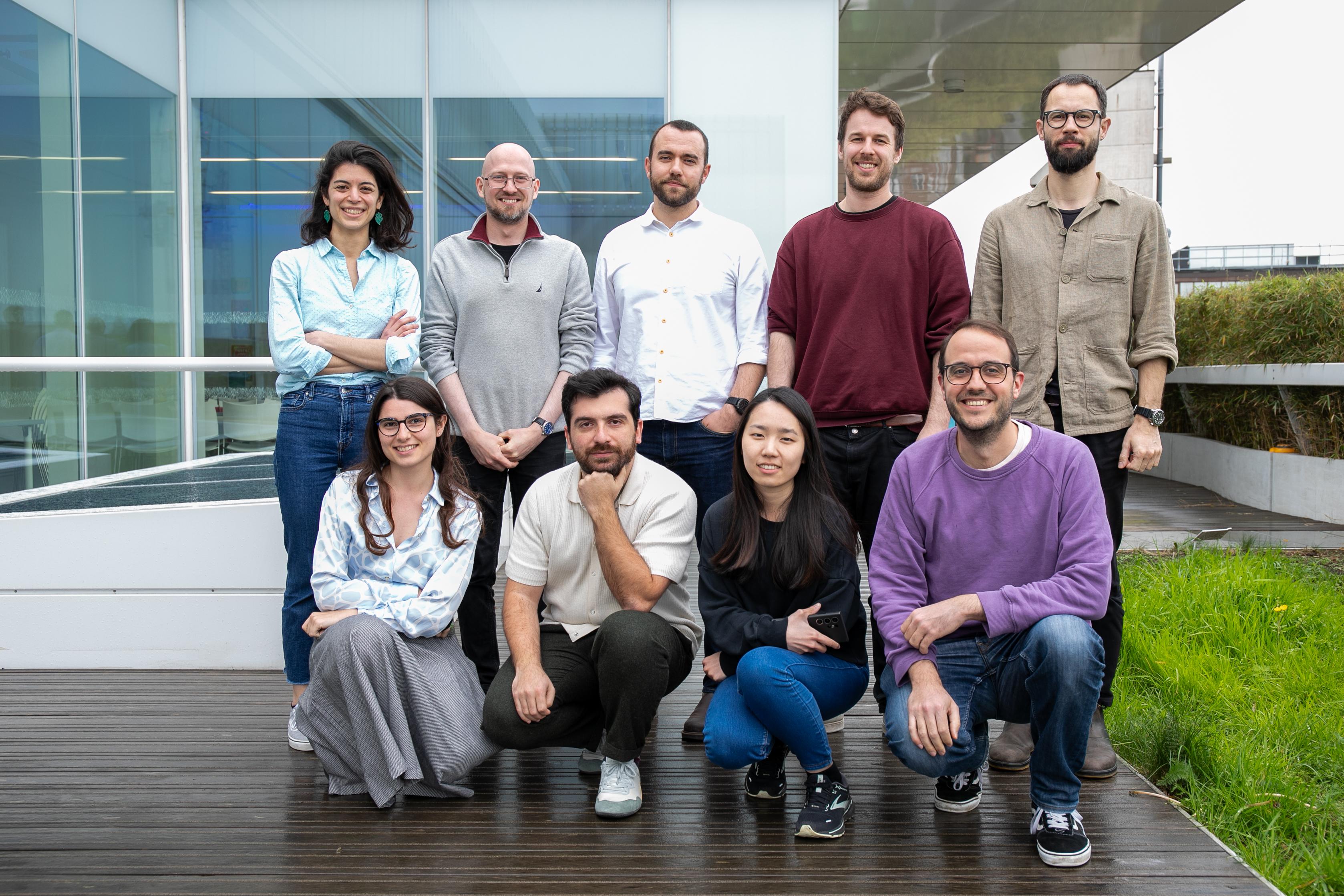

“The idea is to be as public and as accessible as possible with all the work we do. There is no reason why the tools we build should stay private within the institute, particularly because everything we do leverages a rich ecosystem of other open source software,” Adam shared.
Open team science
The various incentives that have driven the traditional model of academic science, as well as experiences with the practicalities of running experiments, informed NIU team members to consider nontraditional career paths in science. “It was an opportunity to stick to parts of the research that I found most interesting, like software pipeline development,” shared Joe Ziminski, when asked about motivations for joining the team. Joe has led recent efforts on developing datashuttle, whose goal is “to standardise the way folders are created for a project that makes data sharing and accessibility easy,” as explained by Laura Porta.
Datashuttle was born out of the need for unifying the way researchers organised their data in the lab. The tool automates the organisation and transfer of experimental data. This simplifies use of this data by others and makes it much easier to build further tools for preprocessing or data analysis. Joe is also working on an electrophysiology pipeline, which will centralise and automate analysis of neural spike data.
Laura joined the NIU to develop the tools neuroscientists need without the pressures of academic publishing. By doing so, she has found meaning in “working and progressing together as a group, bringing something to the community.” This has given her a “different flavour to the work,” emphasising collaboration on shared research objectives over solitary career ambitions. Her work currently ties in closely with projects in the Margrie Lab collaborating with researchers to resolve distortions in rotated microscopy images.
Niko Sirmpilatze shares the view that the progress of a group or investigator’s work tends to remain private until a paper is published. “I think that in science we often claim we do team science. But it’s not always the case because the incentives pull you apart,” he specified. He remarked that how the NIU operates is “very freeing and rewarding because you can work towards something you care about in a team without worrying about who will be the first author or get the grant.”

Alessandro Felder leads the BrainGlobe development efforts at NIU, making the tool easier and more accessible to use by improving graphical user interfaces and focusing on robustness as well as maintainability. He observes that “building general-purpose, easy-to-use software for research contributes to better, more reproducible research”, and notes that part of the research software engineer role includes advocating for more open and collaborative science.
The team operates on such a shared belief that helping science inch toward a better experience for all is a worthy mission. Chang Huan Lo, who focuses on developing behavioural pipelines and polishing website documentation, understands the importance of transparent and accessible data analysis processes for good science. So does Igor Tatarnikov, who works closely with SWC’s Advanced Microscopy Facility. “If we can minimise the amount of effort it takes to go from two-photon microscopy images for neurons firing to analysing data, the more it saves researchers time and effort,” he added.

Addressing obstacles
Adam emphasises how lucky he has been that research software engineering or the general career of building software for science and academia, was founded in the UK and hence is well established. Establishing a team of research software engineers at SWC and GCNU was thus buoyed by the advice of those who came before, both in and outside of UCL. Sofia Miñano-Gonzalez for example, was initially based at UCL’s Advanced Research Computing Centre. “There is a lot less bureaucracy at SWC, so it’s easier to make things happen than it might be in other environments,” Adam remarked.
But challenges were nevertheless present. The perennial problem has been prioritisation. Niko commented, “there are a lot of great enthusiastic scientists who come to us with ideas, and we want to help them. We too are excited about their ideas. But we cannot do everything in parallel, so we have to ask them to wait for their ideas to be executed properly.”
The goals of the NIU team are to build high quality, robust and maintainable tools, and so working on an endless number of projects would not be a strategic move. “We have to be ruthless with prioritisation and focus on building a small number of high quality tools, like movement.”
Enter movement
The movement Python package, released by the team in March, capitalises on the evolving systems neuroscience field’s penchant towards teasing apart the neural bases of animal behaviour. Enabled by software like DeepLabCut or SLEAP, neuroscientists have been able to automatically track animal behaviour in video format, a feat that was hard to imagine even 10 years ago.

“People are conducting sophisticated analyses of tracking each individual point on the body of an animal to extract information about how the animal is behaving,” explained Niko. “But while that part of the analysis is solved – tools that can take a video and locate in every frame the positions of specific body parts – what comes after is not standardised.”
Chang Huan has also worked closely on the movement package. As part of a group of researchers at SWC investigating the neural bases of naturalistic behaviour including foraging, she also dedicates her time to developing and maintaining the application programming interface for SWC’s Project Aeon.
Niko proceeded to point out that users of such software write their own code to import the tracking data into Python, MATLAB, or whichever programming language they use. They also write code to quantify variables such as the speed of the animal, its location in an arena, among others. “What Adam realised is that a lot of the code is repetitive across projects.” When a PhD student or researcher in charge of such a project leaves the lab, for example, the next person is left to start from the beginning and write the same function almost from scratch.
“What we set out to do was identify the common tasks that people do after they’ve completed behavioural tracking and put them all into one Python package that is well designed, maintained, modular and usable across projects,” Niko continued. The movement project was not only inspired by the needs of scientists at SWC, but also prides itself on its usefulness for any behavioural neuroscientist.
Leveraging curiosity
The development of movement from the very beginning was all done publicly on GitHub. All discussions, from to-do lists to meeting minutes, were viewable by everyone around the world. And thanks to this open science ethos of the NIU, the package has received positive feedback from those who were able to witness the preparation and take-off of a versatile toolkit. “We’d still like more feedback on movement, especially from within SWC through talks or seminars,” Niko shared.
Sofia echoed the sentiment. “There is a lot of back-and-forth between us and the users, and you can see that improves the work incrementally,” she noted. “It’s nice to have the chance to work on technical problems in a scientific environment, and create products for questions we feel curious about.”